Trends in Oncology
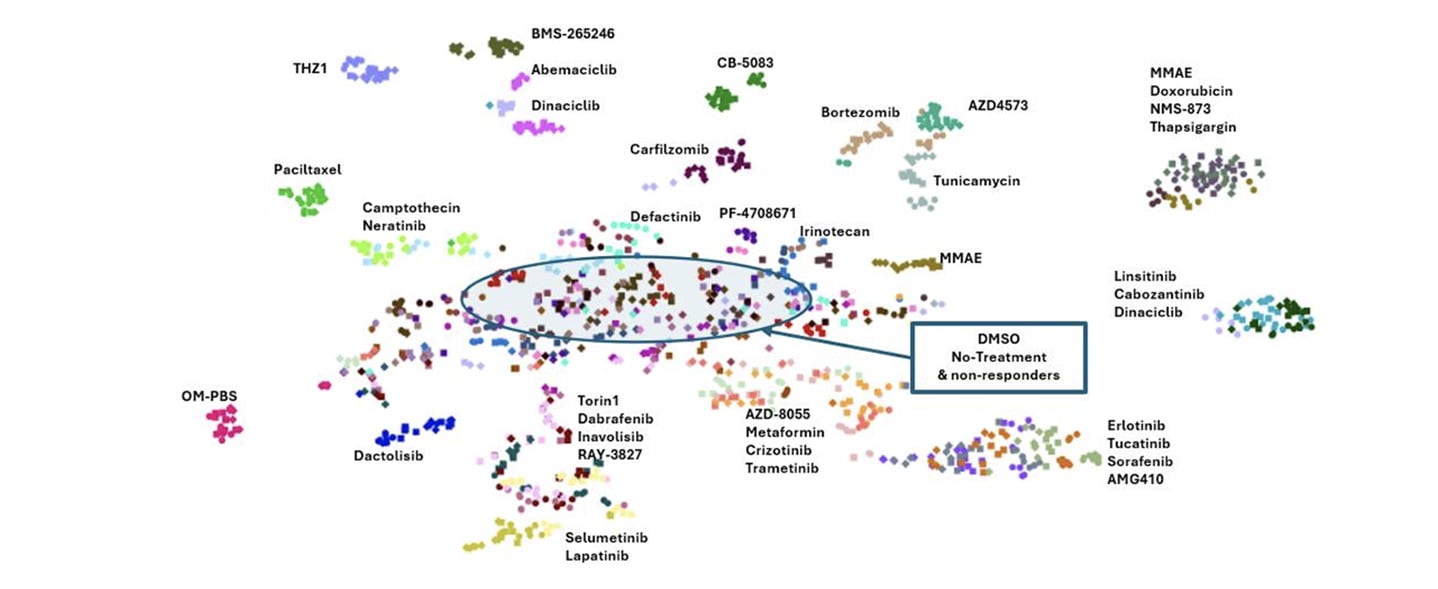
Scaling Chem-Seq and Functional-Seq to Decode Drug and Genetic Perturbations in Patient-Derived Organoids
Large-scale drug discovery increasingly depends on understanding mechanism, not just measuring viability. While high-throughput screening has traditionally relied on simple 2D systems, these models often fail to capture the biological complexity of human tumors or provide the mechanistic depth needed to make confident development decisions. At the same time, transcriptomic approaches have historically been too low-throughput to support large compound libraries. In our AACR 2026 poster, we introduced scalable Chem-Seq and Functional-Seq platforms designed to close this gap. By combining high-throughput transcriptomic profiling with patient-derived organoid models, we can systematically map chemical and genetic perturbations at scale while preserving clinically relevant tumor biology. Why transcriptomic profiling matters in drug discovery Phenotypic screening alone rarely explains why a compound works or fails. Two agents may produce similar viability effects for completely different reasons, while others may appear inactive despite engaging their intended targets. Transcriptomic profiling addresses this challenge by capturing pathway-level responses and signatures of mechanism of action. However, traditional RNA-seq workflows are difficult to scale. Sample preparation, library generation, and sequencing cost all become limiting factors. Our goal was to develop a workflow that preserves transcriptomic resolution while enabling tens of thousands of perturbations per week in relevant cancer models. A scalable Chem-Seq workflow built for throughput Our Chem-Seq platform is built around DRUG-seq™, a workflow optimized for bulk mRNA profiling directly from cancer cell lines or PDX-derived organoids in 384-well plates. Cells or organoids are plated, cultured, and treated with small-molecule compounds. Following treatment, a single-step lysis and barcoded oligo-dT capture allows mRNA from each well to be uniquely indexed in situ. After reverse transcription, all wells are pooled, amplified, and processed through standard Illumina library preparation workflows. Libraries are sequenced to a depth sufficient to capture robust transcriptional signatures while maintaining scalability. This approach supports roughly 50,000 perturbations per week, spanning small molecules, and siRNAs. The key advantage is efficiency. By eliminating many traditional bottlenecks, Chem-Seq enables rapid, reproducible transcriptomic profiling at a scale that aligns with modern screening demands. From raw reads to mechanism-resolved insights Chem-Seq data are processed to generate gene-by-barcode matrices, followed by normalization, batch correction, and dimensionality reduction. Principal component analysis and UMAP visualizations enable rapid triage of perturbation effects, while differential expression and pathway analyses reveal mechanism-associated signatures. To further contextualize results, Chem-Seq signatures are compared against large public perturbation datasets using connectivity mapping approaches. This allows unknown compounds to be matched to known drugs, siRNA knockdowns, or CRISPR perturbations based on shared transcriptional responses, providing early insight into mechanism of action and potential off-target effects. Anchoring chemistry with Functional-Seq While chemical perturbations provide rich information, interpreting their biological meaning is often strengthened by genetic context. Functional-Seq integrates siRNA-mediated knockdown of drug targets directly into the same transcriptomic framework. In this study, Functional-Seq was applied in non-small cell lung cancer PDX organoid models to generate loss-of-function signatures for selected targets. These genetic signatures serve as anchors for interpreting Chem-Seq profiles, enabling direct comparison between pharmacologic inhibition and genetic suppression of the same pathway. Where chemical and genetic perturbations converge transcriptionally, confidence in on-target activity is strengthened. Where they diverge, additional biology, compensatory signaling, or off-target effects can be explored. What the platform reveals in patient-derived organoids Using patient-derived organoids preserves key aspects of tumor heterogeneity and 3D architecture that are often lost in standard cell lines. Across the presented NSCLC PDX organoid models, Chem-Seq generated highly reproducible transcriptional signatures. Compounds targeting shared pathways clustered together, even when structurally distinct, while strongly cytotoxic agents formed separate transcriptional groups. Dose-response profiling added an additional layer of insight. Increasing compound concentration revealed clear transitions between pathway-selective signatures and broader cytotoxic stress responses. This distinction is critical for prioritizing compounds that modulate their intended targets without inducing nonspecific toxicity. Functional-Seq knockdowns recapitulated chemical inhibition for several pathways, demonstrating strong cross-method concordance at the gene and pathway levels. In contrast, targets that failed to produce robust genetic or pharmacologic signatures highlighted limitations related to knockdown efficiency or pathway redundancy. Why integrated Chem-Seq and Functional-Seq matter Together, Chem-Seq and Functional-Seq provide a scalable, mechanism-resolved framework for drug discovery in clinically relevant models. This integrated approach supports: Mechanism confirmation and target engagement assessment Rapid prioritization of compounds with clean, pathway-specific signatures Early identification of off-target or cytotoxic liabilities Improved interpretation of phenotypic screening data Because the platform is compatible with large compound libraries and patient-derived organoids, it bridges the gap between discovery-scale screening and translational relevance. Looking ahead Following validation, this workflow is designed to scale further. Large-scale screening across diverse PDX organoid models and standard-of-care comparators will expand the reference space of transcriptomic signatures. Combined with machine-learning and generative AI approaches, these datasets can be used to predict compound response, guide structure-activity relationships, and even design novel compounds optimized for desired transcriptional outcomes. By integrating high-throughput transcriptomics, genetic perturbation anchoring, and patient-derived 3D biology, Chem-Seq and Functional-Seq offer a powerful foundation for next-generation oncology drug discovery and precision medicine. Want to explore the full data? This blog highlights the overall strategy and key insights, but the complete poster includes detailed workflows, clustering analyses, concordance metrics, and pathway-level results. Download the AACR 2026 poster to explore the full Chem-Seq and Functional-Seq dataset and figures.
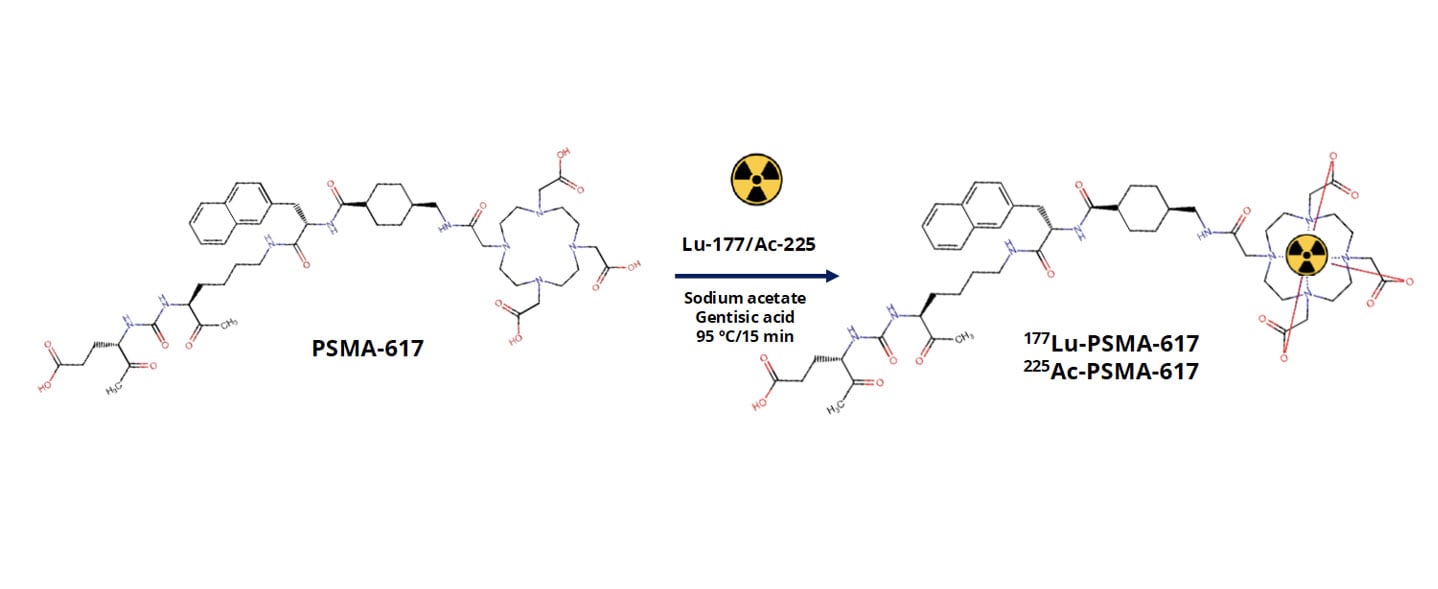
How Preclinical PDX Studies Help Guide PSMA Radioligand Development
Metastatic castration-resistant prostate cancer, or mCRPC, remains a major clinical challenge. While prostate-specific membrane antigen, PSMA, has emerged as a highly promising therapeutic target, differences in tumor biology, target expression, and radiopharmaceutical behavior continue to influence clinical outcomes. For PSMA-targeted radioligand therapies to reach their full potential, developers need preclinical models that reliably capture both tumor complexity and biodistribution behavior in vivo. At AACR 2026, we presented data showing how patientderived xenograft (PDX) models can be used to evaluate PSMA-targeted radiopharmaceuticals at multiple levels. This included radiochemistry quality, biodistribution, and tumor selectivity. The work focused on two clinically relevant agents, [177Lu]LuPSMA617 and [225Ac]AcPSMA617, tested across a panel of mCRPC PDX models with variable PSMA expression. Why PDX models are critical for radiopharmaceutical development Radiopharmaceutical therapies rely on far more than target binding alone. Successful translation depends on tumor uptake, retention, clearance from healthy tissues, and therapeutic index. Traditional cell line models fail to capture the architectural and microenvironmental features that shape these processes in patients. PDX models preserve the genetic heterogeneity and tissue organization of the original human tumor. This makes them particularly well suited for studying radioligand distribution and efficacy in vivo. In the context of PSMA-targeted therapies, PDX models allow us to directly evaluate how differences in PSMA expression and tumor biology influence radiotracer uptake in clinically relevant tumor models. Designing a clinically relevant PSMA radiopharmaceutical study In this study, we evaluated PSMA617 labeled with either lutetium177 or actinium225 across multiple prostate cancer PDX models representing metastatic, castration-resistant disease. The models spanned a range of PSMA expression levels, enabling us to assess how target expression directly relates to biodistribution. We first confirmed robust radiochemical performance. PSMA617 was efficiently labeled with both isotopes, achieving high radiochemical purity that met stringent quality standards. Establishing radiochemistry consistency is a critical first step before moving into in vivo evaluation, particularly for comparative studies involving different isotopes. How biodistribution studies inform target specificity and safety Following radiotracer administration, we conducted biodistribution analyses at defined time points to quantify uptake across tumors and major organs. Rather than relying on a single model, we assessed multiple PDXs to capture biologically driven variability. Across the panel, tumor uptake correlated strongly with PSMA expression levels. Models with high PSMA expression showed selective and robust radiotracer accumulation, while low-PSMA models demonstrated minimal uptake. This clear relationship confirms biological predictability and reinforces the value of molecularly annotated PDX panels when evaluating PSMA-targeted therapies. Importantly, although both [177Lu]LuPSMA617 and [225Ac]AcPSMA617 demonstrated tumor targeting, differences emerged in tumor-to-normal tissue ratios. These distinctions highlight why side-by-side preclinical evaluation is essential when selecting isotopes and dosing strategies for clinical development. Comparing beta- and alpha-emitting PSMA therapies One of the most important questions in PSMA radiopharmaceutical development is how different isotopes behave in vivo. Lutetium177 and actinium225 differ in emission properties, tissue penetration, and potential toxicity profiles. Our comparative biodistribution analyses showed that while both agents localize to PSMA-expressing tumors, [177Lu]LuPSMA617 demonstrated more favorable tumortotissue ratios across several organs. These findings provide early insight into how isotope choice may influence therapeutic window and tolerability. While alpha-emitters remain highly compelling for their potent cytotoxicity, data like this emphasize the importance of rigorous preclinical evaluation in clinically relevant models before advancing treatment strategies. Why this approach matters for radiopharmaceutical pipelines Radiopharmaceutical development is inherently multidisciplinary, integrating chemistry, biology, imaging, and therapeutic evaluation. PDX models provide a unifying platform where all of these components can be assessed in context. Using well-characterized mCRPC PDXs allows us to: Link PSMA expression directly to in vivo uptake Compare isotopes in clinically relevant tumor models Evaluate tumor selectivity alongside therapeutic response Reduce translational uncertainty before clinical studies This kind of integrated preclinical strategy is especially valuable as PSMA-targeted therapies continue to expand into earlier disease settings and combination regimens. Want to see the full dataset? This blog highlights key findings, but the full study includes detailed radiochemistry validation, biodistribution data, tumor-to-tissue ratios, and efficacy results across multiple PDX models. Download our AACR 2026 poster to explore the complete results and analyses.
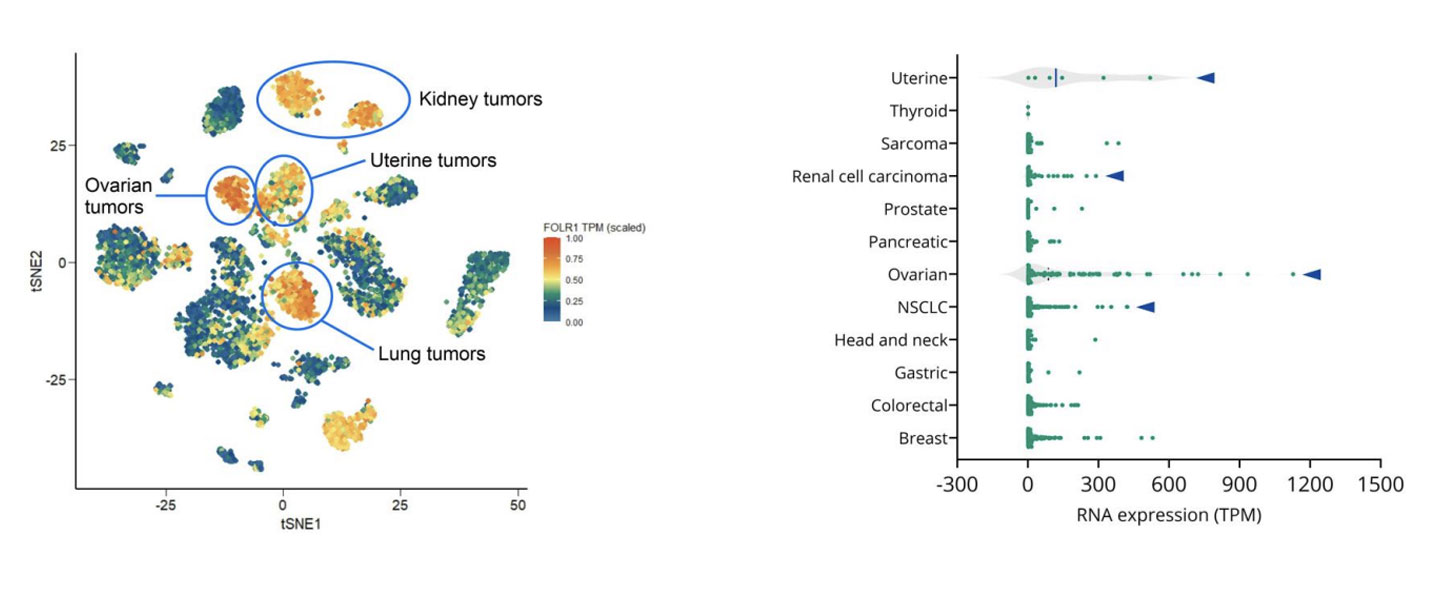
How to Use PDX Models to Advance Folate Receptor-Alpha ADC Development in Ovarian Cancer
Epithelial ovarian cancer remains one of the deadliest gynecologic malignancies. Although many patients initially respond well to platinum-based chemotherapy, relapse is common, and outcomes for platinum-resistant disease have seen little improvement over the past decade. With limited treatment options and a shortage of robust predictive biomarkers, there is a clear need for more precise therapeutic strategies that can better match patients to targeted therapies. One promising area of progress is antibody-drug conjugates, or ADCs. These therapies are designed to deliver potent cytotoxic agents directly to tumor cells by targeting tumorassociated antigens. In ovarian cancer, folate receptoralpha, or FRα, has emerged as an especially compelling target due to its high expression in ovarian tumors and comparatively limited expression in normal tissues. At AACR 2026, we shared new data showing how patient-derived xenograft, or PDX, models can be used to evaluate FRα-targeted ADCs, explore variability in response, and begin to uncover potential mechanisms of resistance. This work highlights how clinically relevant PDX models, combined with molecular profiling, can support smarter ADC development decisions earlier in the pipeline. Why FRα is an attractive ADC target in ovarian cancer FRα is a transmembrane glycoprotein involved in folate transport and cellular metabolism. While healthy tissues show limited FRα expression, ovarian tumors, particularly high-grade serous carcinomas, often express it at high levels. This contrast makes FRα well suited for ADC approaches that aim to maximize tumor-specific activity while minimizing offtarget toxicity. Mirvetuximab soravtansine-gynx, marketed as Elahere®, is an FRα-targeted ADC recently approved for patients with FRα-positive, platinum-resistant ovarian cancer. However, clinical experience has shown that FRα expression alone does not reliably predict which patients will respond to treatment. Understanding this response heterogeneity is one of the key challenges in optimizing ADC development. How we select the right PDX models to reflect patient biology To address this challenge, we leveraged our gynecologic cancer PDX collection, which includes models derived from both treatment-naïve and heavily pretreated ovarian cancer patients. Using integrated RNA sequencing and immunohistochemistry, we assessed FRα expression across ovarian PDXs and compared these patterns with patient data. Consistent with clinical observations, ovarian cancer PDXs showed some of the highest FOLR1 expression levels across tumor types. Importantly, RNAbased expression data strongly correlated with membrane-specific FRα protein expression. This confirms that multiple complementary methods can be used to select biologically appropriate models for FRαtargeted ADC screening. Rather than relying on a single cutoff, this approach allows us to build PDX cohorts that better reflect the biological diversity seen in the clinic. What PDX efficacy studies reveal about ADC response Selected ovarian PDX models were treated with mirvetuximab soravtansine using a clinically relevant dosing schedule. As expected, models lacking FRα expression did not respond to treatment, reinforcing the importance of FRα as a prerequisite for activity. Among FRα-positive models, however, responses varied widely. Some tumors showed meaningful growth inhibition, while others demonstrated limited or no response despite high FRα expression. This mirrors what is seen in the clinic, where only a subset of FRαpositive patients achieves durable benefit from treatment. These results highlight a critical lesson for ADC development. Target expression is necessary, but it is rarely sufficient on its own. Using molecular analysis to explore resistance mechanisms To begin understanding why FRα-positive tumors respond differently, we compared responding and non-responding PDX models using transcriptomic and pathway-level analyses. Rather than focusing on single genes, we looked at broader biological processes that may influence sensitivity or resistance to ADC therapy. Clear differences emerged between sensitive and resistant tumors. In particular, pathways related to intracellular transport and drug efflux were enriched in non-responding models. Several of these pathways involve ATP-binding cassette transporters, which are well known for their role in multidrug resistance. While exploratory, these findings suggest potential mechanisms by which tumor cells may evade ADC-delivered payloads, even when the target antigen is present. This type of insight can help guide biomarker development and inform rational combination strategies. How we model acquired resistance using PDXs Primary resistance is only part of the challenge. In many patients, response to ADCs is followed by progression under treatment. To study this, we are actively generating PDX models from patients who initially responded to ADC therapy and later relapsed. In the AACR study, one such PDX was derived from a patient who had responded to mirvetuximab soravtansine before progressing. When re-challenged in vivo, this model showed resistance to treatment, suggesting that clinically acquired resistance mechanisms can be preserved in PDXs. Models like this provide powerful tools for studying resistance biology and evaluating nextgeneration therapies designed to overcome it. Why PDXs matter for ADC development Taken together, these findings show how PDX models can support ADC development beyond basic efficacy screening. When combined with molecular and bioinformatics analyses, they can help: Improve patient stratification beyond single biomarkers Reveal biological drivers of heterogeneous response Enable investigation of intrinsic and acquired resistance Support the development of next-generation ADCs and combination approaches As ADC pipelines continue to expand, integrating clinically representative PDX models early in development can improve translation and, ultimately, patient outcomes. Want to explore the full data? This blog highlights key insights, but the full dataset includes detailed efficacy results, pathway analyses, and figures that provide additional depth. Download our AACR 2026 poster to explore the complete results and analyses.
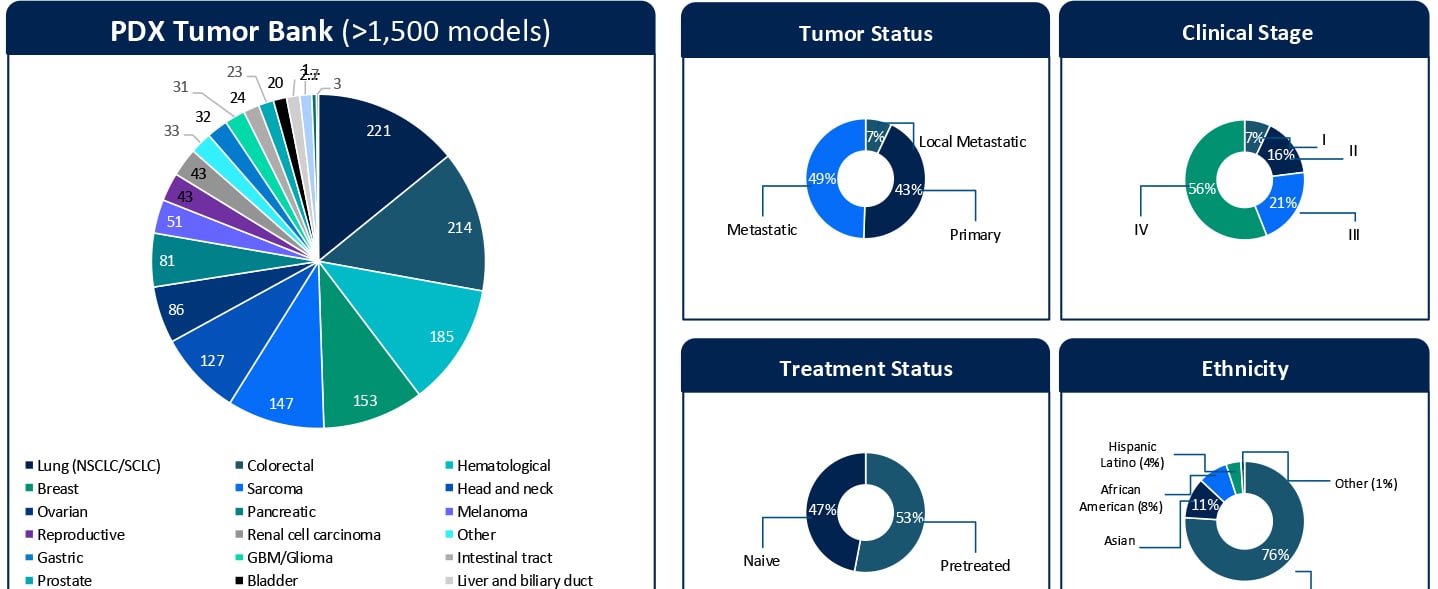
Understanding Sensitivity and Resistance to Pan-RAS(ON) Inhibition Across KRAS-Mutant Tumors
Understanding Sensitivity and Resistance to Pan-RAS(ON) Inhibition Across KRAS-Mutant Tumors KRAStargeted therapies are entering a new phase of clinical relevance. Daraxonrasib, a next-generation pan-RAS(ON) inhibitor designed to target a broad spectrum of KRAS G12X mutations, has shown impressive clinical activity. In a recent Phase III RASolute 302 trial for second-line metastatic pancreatic ductal adenocarcinoma (PDAC), daraxonrasib (RMC-6236) nearly doubled the median overall survival for patients to approximately 13.2 months, compared with 6.7 months for standard chemotherapy.1 These results highlight the rapid progress in this space. At the same time, clinical development is expanding KRAS-targeted therapies into earlier lines of therapy. Daraxonrasib is currently being evaluated in first-line settings, both as monotherapy and in combination approaches. While both clinical and preclinical data demonstrate activity across multiple tumor types, variability in response and the emergence of resistance remain key challenges that limit durable benefit. Additionally, the molecular determinants that define sensitivity, primary resistance, and acquired resistance remain incompletely understood. Addressing these gaps in our understanding of the molecular drivers of sensitivity and resistance is critical not only for patient selection but also for informing rational combination and sequencing strategies in an evolving treatment landscape. A Translational Platform to Study KRAS Biology in Context To better understand these response patterns, this study leveraged Champions Oncology’s low-passage, clinically annotated Patient-Derived Xenograft (PDX) Tumor Bank. With more than 1,500 models and extensive molecular and treatment history annotation, this platform captures both tumor heterogeneity and clinically relevant treatment context. Importantly, approximately half of the models represent pretreated disease, reflecting real-world patient populations. The translational relevance of this platform is supported by prior clinical correlation analyses performed using Champions’ low-passage PDX models. In a study by Izumchenko et al., treatment responses observed in these models were directly compared with clinical outcomes in matched patients across 92 individuals and 129 therapies.2 Strong concordance between the PDX and patient response was observed, with a positive predictive value of approximately 85% and a negative predictive value of approximately 91%. These findings establish Champions’ low-passage PDX models both as a tool for biological exploration and a clinically validated system for informing translational and drug development decisions. In this study, more than 50 KRAS-mutant PDX models were screened across non-small cell lung cancer, colorectal cancer, and pancreatic cancer (PDAC). The cohort included diverse KRAS alleles, co-occurring genomic alterations, and varying treatment histories, which allowed treatment response to be evaluated within a realistic molecular and clinical framework. Heterogeneous Responses Reflect Tumor-Specific Context Daraxonrasib treatment resulted in a broad spectrum of responses, ranging from strong disease regressions to progressive disease. Response variability was not solely explained by KRAS mutation status alone but was influenced by the broader genomic context of each tumor. For example, alterations such as MYC amplification were observed in both responding and non-responding models, suggesting that primary resistance is context-dependent and likely driven by multiple interacting genomic factors, rather than single genomic features. These findings highlight the importance of distinguishing between intrinsic resistance mechanisms and those that emerge under therapeutic pressure. Probing Primary Resistance in KRAS G12D Pancreatic Models Given the clinical relevance of KRAS G12D in pancreatic cancer, a focused analysis was performed in this subset. Differential gene expression analysis comparing responders and non-responders identified candidate markers associated with response, including MSLN and CYP24A1. While these findings are exploratory and based on a limited sample set, they provide an initial framework for stratifying tumors and generating hypotheses around mechanisms of intrinsic resistance. Ongoing work is expanding these analyses through deeper multi-omic integration, including whole exome sequencing (WES), RNA sequencing, proteomics, phosphoproteomics, and surface proteomics. These datasets will be interpreted in the context of Champions’ clinically annotated models, which include detailed treatment histories, to better understand the biological drivers of response and resistance and to support more robust biomarker development. Modeling Acquired Resistance Under Continuous Drug Pressure Selected PDX models were treated continuously beyond their initial response to investigate how resistance evolves over time. A particularly illustrative example is model CTG-2473, a KRAS G12D pancreatic cancer model derived from a tumor collected after first-line gemcitabine treatment. This model reflects a clinically relevant treatment sequence where patients, following standard chemotherapy, would be transitioned to KRAS-targeted therapies such as daraxonrasib. Therefore, treatment of CTG-2473 with daraxonrasib can model response and resistance in a post-chemotherapy setting that mirrors emerging clinical practice. Under continuous treatment pressure, this model demonstrated initial tumor regression followed by relapse, which is consistent with the development of acquired resistance observed in patients. This creates a powerful framework for translational studies. Experiments can be designed in the parental setting to evaluate therapeutic switching upon relapseto assess next-line agents or rational combination strategies. In parallel, Champions Oncology is actively developing a fully resistant subline through serial passaging under sustained drug pressure. This will enable deeper molecular characterization of acquired resistance mechanisms and provide a robust system for testing strategies for overcoming resistance. Together, these models establish a framework not only to observe resistance but also to actively model clinically relevant treatment sequencing scenarios. Why Clinically Annotated PDX Models Matter A defining strength of this work is the integration of molecular profiling, treatment response, and detailed clinical annotation within a single experimental system. Unlike conventional preclinical models, Champions’ PDX platform enables analysis of tumor behavior in the context of prior therapies, disease progression, and patient-specific treatment history. This is particularly important for targeted therapies such as KRAS inhibitors, where response and resistance are shaped by both molecular context and prior treatment. When combined with multi-omic datasets, including WES, RNA sequencing, proteomics, phosphoproteomics, and surface proteomics, these clinically annotated models provide a systems-level understanding of tumor biology. This supports the identification of candidate biomarkers and the development of mechanistically informed combination and sequencing strategies. Importantly, this integrated approach allows researchers to move beyond static response measurements tto capture how tumors dynamically adapt under therapeutic pressure and how these adaptations can be therapeutically targeted. From Mechanism to Strategy Taken together, these data demonstrate that response to pan-RAS(ON) inhibition is shaped by tumor-specific molecular context and dynamic adaptation under treatment. Understanding both primary and acquired resistance will be essential for translating KRAS-targeted therapies into durable clinical benefit. The full dataset, including detailed response profiles, genomic annotations, and resistance modeling, is presented in the accompanying poster. Download the Full Poster To explore the complete results, figures, and molecular analyses, including responder and non-responder comparisons and acquired resistance models, download the full poster here. References Daraxonrasib Demonstrates Unprecedented Overall Survival Benefit in Pivotal Phase 3 RASolute 302 Clinical Trial in Patients with Metastatic Pancreatic Cancer | Revolution Medicines [Internet]. Revolution Medicines. 2026. Available from: https://ir.revmed.com/news-releases/news-release-details/daraxonrasib-demonstrates-unprecedented-overall-survival-benefit Izumchenko E, Paz K, Ciznadija D, Sloma I, Katz A, Vasquez-Dunddel D, et al. Patient-derived xenografts effectively capture responses to oncology therapy in a heterogeneous cohort of patients with solid tumors. Annals of Oncology. 2017 Oct;28(10):2595–605.
Clinically Relevant 3D Tumor Models for HER2-Targeted ADC Development
Expanding the Preclinical Toolbox with a Novel Patient-Derived Organoid Platform The human epidermal growth factor receptor 2 (HER2) plays a central role in regulating cell growth, division, and survival. When overexpressed, it acts as an "on switch", driving accelerated cell growth and uncontrolled tumor spread. This biology underlies the aggressive behavior observed in HER2-positive cancers, which is associated with higher recurrence rates and poorer patient outcomes. While genomic changes in HER2 are most frequently discussed in the context of breast cancer, HER2 alterations are also found in non-small cell lung, ovarian, colorectal, and pancreatic cancers. HER2-Targeted ADCs in Cancer Therapy HER2-targeted therapies have significantly changed the treatment landscape and brought new hope and better outcomes for patients. Among them, HER2-targeted antibody-drug conjugates (ADCs) have emerged as a particularly promising approach, combining the specificity of antibodies with the potency of cytotoxic payloads. These agents enable targeted delivery directly to HER2-expressing tumor cells while limiting off-target toxicity. FDA approved drugs in this class include trastuzumab emtansine (T-DM1, Kadcyla®) and trastuzumab deruxtecan (T-DXd, Enhertu®), both of which have demonstrated meaningful clinical responses (Figure 1). A growing number of next-generation HER2 ADCs are now in development. Figure 1. FDA approved HER2-targeted ADCs trastuzumab emtansine (T-DM1, Kadcyla®) and trastuzumab deruxtecan (T-DXd, Enhertu®). Figure adapted from Joubert et al1. As ADCs become an increasingly important therapeutic modality, the limitations of conventional preclinical models have become magnified. Drug developers need better, more physiologically relevant and predictive preclinical models to support ADC development, ones capable of reproducing tumor architecture, heterogeneity, and treatment response. Limitations of Traditional 2D Preclinical Models for ADC Evaluation Despite their widespread use, traditional 2D cell culture models have well-recognized drawbacks with respect to evaluating targeted therapies. They often fail to capture key biological features of tumors and inadequately replicate the complex tumor microenvironment (TME) that influences drug response in patients. Target accessibility, receptor density, internalization kinetics, and payload penetration are all influenced by three-dimensional tumor structure, features which are largely absent in 2D cultures. As a result, 2D assays may overestimate drug potency or fail to distinguish on-target activity from nonspecific cytotoxicity, reducing their predictive value for clinical translation. In contrast, growing tumor cells as 3D organoids allows for a spatially organized structure that more closely resembles the in vivo microenvironment and architecture of patient tumors. Champions has developed a scalable 3D screening platform based on HER2-positive patient-derived xenograft organoids (PDXOs). Derived from well-characterized PDX models, our ex vivo 3D organoids maintain clinically translatable HER2 expression, providing a relevant and predictive platform for evaluating the efficacy of HER2-targeted ADCs. Validation of HER2-Positive PDXO Models HER2-positive ex vivo PDXO breast cancer models generated from our TumorGraft3D (CTG-3D) platform were evaluated for HER2 expression as well as functional responses to the clinically approved HER2-targeted ADCs, trastuzumab emtansine (T-DM1) and trastuzumab deruxtecan (T-Dxd). The workflow is shown in Figure 2. Figure 2. Workflow overview to evaluate HER2-targeted ADCs using breast cancer PDXO models. HER2 Expression Immunohistochemical (IHC) analysis of PDXO tissue blocks demonstrated complete and intense membrane staining (HER2 score 3+) across HER2-positive models. Consistent with these findings, flow cytometric analysis of enzyme-dissociated organoids identified HER2-positivity in 61.8% to 90.1% of cells. In contrast, the HER2-negative control PDXO model showed no detectable HER2 staining by IHC and negligible HER2 expression by flow cytometry, confirming assay specificity and the absence of nonspecific background signal. A subset of the results is shown in Figure 3. Figure 3. HER2 expression assessed by IHC and flow cytometry analysis. A) HER2-positive PDXO and B) HER2-negative PDXO model. Together, the IHC staining and flow cytometry results are consistent with the clinical annotation and indicate that HER2 expression is maintained relatively homogeneously in breast cancer PDXO models. Response to HER2-Targeted ADCs After confirmation of HER2 expression, the PDXO models were evaluated for their response to two clinically approved HER2-targeted ADCs, trastuzumab deruxtecan (T-DXd) and trastuzumab emtansine (T-DM1). Both ADCs were tested against HER2-positive and HER2-negative breast cancer PDXOs and reference breast cancer cell lines. In addition to intact ADCs, corresponding cytotoxic payloads (Exatecan and DM1), naked antibody (trastuzumab), and—for T-DXd—an isotype-payload conjugate (IgG-DXd) were evaluated to distinguish potency, selectivity, and target-dependent activity. Drug response was measured using the CellTiter-Glo® viability assay following six days of incubation with the ADCs and control treatments. Figure 4. Representative dose-response curves (IC₅₀). A) HER2-positive PDXO model comparing exatecan (payload) and trastuzumab deruxtecan (T-DXd) and B) control naked antibody trastuzumab and isotype-payload conjugate IgG-DXd. Across HER2-positive breast cancer PDXO models and control cell lines, ADCs demonstrated greater efficacy and specificity compared with naked antibody and isotype-payload controls (Figure 4, and data not shown). Full datasets across additional PDXO models are presented in the accompanying AACR 2025 poster. These results highlight the utility of PDXO models derived from Champions’ CTG-3D platform for evaluating next-generation ADCs and discriminating between payload-driven cytotoxicity, nonspecific conjugate effects, and HER2-targeted ADC activity. To further quantify differential treatment responses, normalized Area Under the Curve (∆AUC) values were calculated to capture response differences between isotype controls and ADCs across the full concentration range. HER2-positive breast cancer cell lines and PDXO models exhibited higher ∆AUC values (approximately 30–50%, data not shown), indicating stronger ADC activity relative to controls. In contrast, HER2-negative models showed substantially lower ∆AUC values (~20%), consistent with reduced ADC activity in the absence of target expression (Figure 5). Figure 5. Normalized ∆AUC. The response difference between isotypes and ADCs is calculated as: [AUC of Isotype] – [AUC of ADC] / [AUC of Isotype ] x 100% Collectively, these findings demonstrate that breast cancer PDXO models generated from Champions’ CTG-3D platform reproduce expected response patterns to FDA-approved HER2-targeted ADCs with known pharmacological profiles, supporting their utility as a predictive ex vivo platform for ADC development. Key Takeaways Champions’ CTG-3D platform is a biologically relevant ex vivo 3D platform that can support critical preclinical decisions, including payload comparison and mechanism-based screening, enabling ADC candidates to be screened and prioritized before more expensive in vivo evaluation and clinical development is undertaken. For drug developers, this translates into greater confidence in candidate selection, improved translational relevance of preclinical data, and reduced risk of late-stage attrition of potentially viable candidates. By enabling better-informed decisions earlier in the development process, the CTG-3D platform helps accelerate the advancement of safer and more effective ADC-based therapies. To explore the full datasets in more detail, download the full poster presented at AACR 2025 here. References Joubert, Nicolas, Alain Beck, Charles Dumontet, and Caroline Denevault-Sabourin. 2020. “Antibody-Drug Conjugates: The Last Decade.” Pharmaceuticals (Basel, Switzerland) 13 (9). https://doi.org/10.3390/ph13090245.
Evaluating a Novel GCN2 Inhibitor for Acute Myeloid Leukemia (AML) Treatment Using Patient-Derived Models
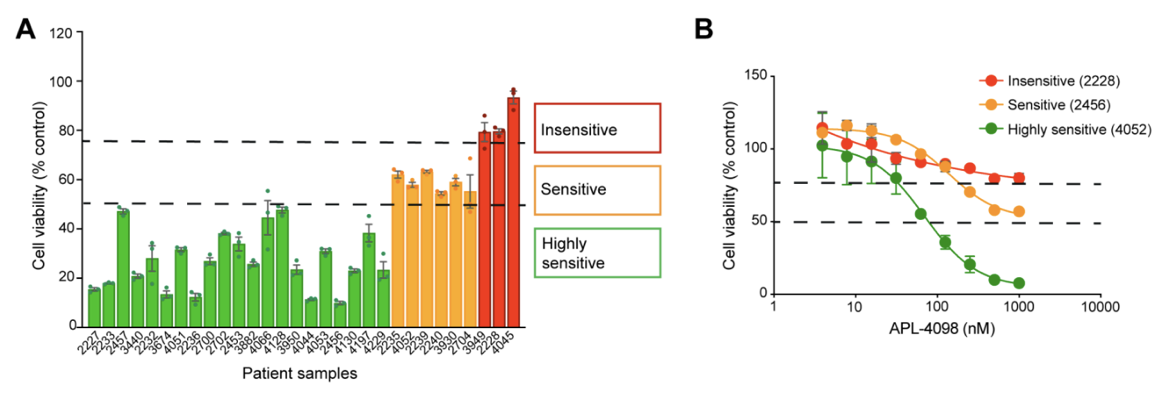
The AML Therapeutic Landscape Relapse remains one of the greatest challenges in treating Acute Myeloid Leukemia (AML). While targeted therapies have improved outcomes for some patients, high relapse rates and limited durable treatment options mean that many patients still face poor prognoses. One pathway gaining attention is the Integrated Stress Response (ISR), a highly conserved protective signaling network that helps cells adapt to nutrient deprivation, oxidative stress, and other metabolic stressors. Many tumor types, including AML, depend heavily on this stress-response system to survive and proliferate. Within this pathway, the kinase GCN2 (General Control Nonderepressible 2) serves as a key sensor of amino acid deprivation and metabolic stress, helping cancer cells survive under unfavorable conditions where lack of exogenous nutrients may disturb intracellular homeostatic balance. Disrupting this survival mechanism has emerged as a promising strategy to selectively target cancer cells and improve treatment outcomes. A recent study explored a novel and selective inhibitor of the stress-response kinase GCN2, APL-4098, as a potential therapeutic approach for AML1. Ex Vivo Evaluation of APL-4098 in Patient-Derived AML Cells Román-Trufero et al. first characterized the anti-leukemic activity of APL-4098 in ex vivo primary patient AML samples using Champions Oncology’s AML VitroScreen platform. A cohort of 30 patient-derived samples was treated with increasing concentrations of the compound and cell viability was measured using a luminescence-based viability readout to determine how effectively the drug inhibited leukemia cell growth. From these results, dose–response curves and IC50 values were generated to quantify drug potency across the cohort. The results revealed strong inhibitory activity across the patient cohort with 70% of samples (21/30) showing a ≥50% reduction in cell viability, indicating high sensitivity to the GCN2 inhibitor. An additional 20% of samples demonstrated some sensitivity, with a 25–50% reduction in viability. Finally, 10% of the samples showed less than 25% decrease in viability and were categorized as insensitive (Figure 1). Figure 1. APL-4098 cytotoxic activity ex vivo. (A) Effect of 500 nM APL-4098 on the viability of patient-derived AML samples after 72h treatment. Values for each specimen are expressed relative to each sample’s matched vehicle-treated control (normalized to 100%) and presented as mean ± SEM, n=3. (B) Selected APL-4098 dose-response curves of highly sensitive (green), sensitive (orange), and insensitive (red) patient-derived samples. Notably, responses were observed across diverse genomic backgrounds, with no clear correlation between common AML mutations and sensitivity. Cells from pre-treated or relapsed patients also responded, suggesting that GCN2 inhibition may be effective even in difficult-to-treat AML. This ex vivo screening step provided critical insight into which patient samples were most sensitive, laying the groundwork for subsequent in vivo evaluation in patient-derived xenograft (PDX) models. In Vivo Evaluation of APL-4098 in AML Patient-Derived Xenografts Following ex vivo screening, the anti-leukemic activity of APL-4098 was further assessed in Champions’ patient-derived xenograft (PDX) AML models. Human AML cells from a patient sample that showed ex vivo sensitivity to APL-4098 were implanted into immunodeficient NOG mice via tail vein injection, allowing leukemia to engraft in the bone marrow. Engraftment of human AML cells in the bone marrow was monitored in surrogate animals using flow cytometry, and once engraftment reached ~20% human CD45+ cells, mice were randomized into four treatment groups: Vehicle control (DMSO) APL-4098 alone (15 mg/kg once daily) Venetoclax alone (100 mg/kg once daily), as a standard-of-care comparator APL-4098 + venetoclax combination Treatments were administered daily for 19 days, and leukemia burden was evaluated by analyzing multiple AML subpopulations, including blasts, progenitors, and leukemia stem cells (LSCs), using multicolor flow cytometry (Figure 2). This allowed researchers to evaluate both single-agent activity and potential synergy with venetoclax in a clinically relevant AML model. Figure 2. Effect of APL-4098, Venetoclax and the combination of both in vivo in an AML PDX model. Bone marrow from AML-engrafted animals was assessed for (A) the percentage of viable human CD45+. (B) the number of viable blasts (CD34+/CD33+/-). (C) the number of viable AML progenitors (CD34+/CD38+) (D) the number of viable LCSs (CD34+/CD38-). In all graphs, data shown as mean ± SEM, statistical significance determined by one-way ANOVA. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001. DMSO, n=6; APL-4098, n=7; Venetoclax, n=8, APL-4098 + Venetoclax, n=7. The results showed APL-4098 as a monotherapy had little to no effect on bulk leukemic burden, but preferentially targeted and reduced the LSC-enriched compartment, a niche generally considered primarily responsible for relapse initiation after bulk eradication of leukemic blasts2 Importantly, the combination of APL-4098 and Venetoclax, the standard of care BCL-2 inhibitor in AML patients, produced a pronounced synergistic response, achieving a 97% to 99% reduction across all leukemia subpopulations analysed. Mechanistic studies, including RNA sequencing and metabolic analyses, suggested that APL-4098 exerts its anti-leukemic activity by suppressing mitochondrial respiration, triggering the mitochondrial unfolded protein response, and inducing metabolic stress in AML cells. Together, the researchers demonstrated APL-4098 effectively induces cytotoxicity both ex vivo and in vivo, with preferential effect against the LSC subpopulation, a key driver of patient relapse. While APL-4098 shows potential as a monotherapy agent in the preclinical setting, the potent synergistic interaction between APL-4098 and Venetoclax highlights a potential combination therapy for AML that merits further non-clinical and clinical exploration. This study demonstrates the utility of Champions Oncology’s patient-derived platforms for preclinical and translational AML research, providing a framework to evaluate drug efficacy and explore new therapeutic strategies. To learn more about this study, download the full publication here Explore the Champions Oncology website to discover more about our hematological cancer models and testing capabilities References Román-Trufero M, Whitlock G, Seydoux C, et al. A novel potent and selective GCN2 inhibitor, APL-4098, has anti-leukemic activity through dysregulation of mitochondrial function. Clin Cancer Res. 2026. Hansen Q, Bachas C, Smit L, Cloos J. Characteristics of leukemic stem cells in acute leukemia and potential targeted therapies for their specific eradication. Cancer Drug Resist. 2022.
Not All Tumors Look Alike: Using Surfaceome Diversity to Enrich Responders
The case for measuring the surface directly Biologics and antibody drug conjugates operate at the cell's surface, yet many programs still guide patient selection using RNA expression or whole cell protein abundance. That practice can misclassify candidates because transcripts and total protein often do not correlate, due to post-transcriptional and post-translational mechanisms. Moreover, cellular localization and receptor density can’t be quantified by using whole cell bulk proteomics, therefore not guaranteeing that a receptor is exposed on the exterior membrane above drug relevant thresholds. In 2024, investigators profiled the surfaceome of 100 genetically diverse, primary human AML specimens and resolved antigen patterns on primitive and stem like cells with limited expression in essential normal tissues. The work demonstrated that surface level heterogeneity is real, clinically meaningful, and different from what one would infer from bulk RNA or whole cell proteomics alone. Cell+1 Calls to better map the surfaceome are growing. A 2024 Cancer Discovery commentary noted that only a few dozen cell surface targets currently anchor FDA or EMA approved therapies, despite the surface being the most accessible compartment for antibodies, CAR T cells, and radiopharmaceuticals. The authors argued for intensified efforts to chart the universe of surface proteins across cancers, which would accelerate target discovery and improve translational relevance for modalities that require true surface accessibility. AACR Journals+1 Solving the Surfaceome Problem Why a surface resolved view changes development plans Modern reviews describe how advances in mass spectrometry, fractionation, and enrichment are making it feasible to survey the cancer surfaceome at scale. These approaches improve the identification of drug accessible proteins that whole cell proteomics can miss, because abundant intracellular proteins dominate unfractionated measurements and mask low abundance, surface localized receptors. The reviews also outline practical ways to mitigate technical challenges such as low copy number, hydrophobic domains, and contamination by intracellular compartments, for example through careful membrane preparation, parallel intracellular and whole cell fractions, and stringent enrichment and quality control thresholds. The program level implication is straightforward. When a therapy acts at the surface, the biomarker strategy should resolve surface localization directly rather than infer it from RNA or whole cell protein alone. PubMed+1 A second implication concerns internalization and trafficking. For ADCs and some bispecifics, internalization kinetics and routing to lysosomes affect payload delivery and potency. Surface resolved workflows can be coupled to orthogonal assays, for example flow cytometry with ligand induced internalization or live cell imaging, to determine whether a receptor is not only present at the surface but also behaves in a way that supports the intended mechanism. This behavioral dimension rarely appears in transcript based selection but influences both efficacy and safety. From heterogeneity to practical enrichment Turning surface heterogeneity into clinical signal requires a sequence of disciplined steps. First, measure what matters. Use fractionated surface proteomics or validated surface specific immunohistochemistry and flow assays that distinguish the exterior membrane from total abundance. Second, define actionable thresholds that tie expression to benefit and that can be reproduced across sites. Educational content from the ASCO Educational Book emphasizes that quantitative cutoffs and standardized assays are central to patient selection for antibody drug conjugates, because target accessibility and abundance determine benefit, and because inconsistent thresholds erode the interpretability of early phase studies. Third, map prevalence by histology and by clinically relevant subgroups, since an attractive target with low prevalence may not support enrollment or may require a targeted site strategy. When these elements are present, enrichment reflects how the drug actually works and not a proxy signal. ASCO Publications+1 Surface level diversity can also sharpen indication strategy. If a receptor is highly enriched at the surface in a subset of colorectal cancer but not in pancreatic cancer, a program can prioritize the former for first in human evaluation, even if RNA levels are similar in both. This is particularly relevant where surface and whole cell abundances diverge. The AML study noted above showed precisely this phenomenon, where antigen exposure on stem like compartments could be quantified and connected to therapeutic concepts that require surface binding. Cell Selectivity and safety as design constraints Surface heterogeneity intersects with safety because many candidate antigens have some expression in normal tissues. A 2024 review in Trends in Pharmacological Sciences surveyed strategies that increase antibody selectivity in oncology, including superselectivity through avidity and multivalency, conditional or pH sensitive binding, dual targeting that requires co expression to achieve high affinity, and engineering designs that leverage tissue context. These concepts translate directly into earlier program decisions. Targets that can be paired with selective engineering approaches should score higher than those that would require unrealistic discrimination from a conventional binder. A transparent scoring framework, for example one that penalizes normal tissue expression in essential organs and rewards tumor specific co expression patterns, makes those choices auditable during governance reviews. Cell+1 The safety argument is not theoretical. ADCs can cause off tumor toxicities when a payload is delivered to normal tissues that express the target at modest levels. A biomarker plan that quantifies true surface exposure in disease and screens for surface exposure in a curated panel of normal tissues improves the chance of a workable therapeutic window. Reviews that focus on ADC biomarkers, together with trial design guidance, point in the same direction, namely that selection should be quantitative and assayable, not an exploratory cutpoint chosen after the fact. ASCO Publications Concrete examples of surfaceome driven discovery Evidence that surface resolved discovery translates into programs is accumulating. In late 2024, a Cancer Cell study used an integrative proteogenomic surfaceome approach to credential DLK1 as an immunotherapeutic target in neuroblastoma. The authors combined mass spectrometry based surface profiling with genomic and transcriptomic context to demonstrate surface exposure, tumor specificity, and functional relevance, then moved to validation experiments that supported drug development. This work illustrates how surface level datasets, anchored in clinical material, can identify and qualify targets that a transcript only screen might underrate. PMC+3Cell+3PubMed+3 Surfaceome maps are also becoming more accessible, which will help teams generalize findings across institutions. The AML dataset is public in GEO, enabling independent inspection of QC criteria, antigen lists, and statistical methods. Commentary from Cancer Discovery underscores the systemic opportunity, arguing that better cartography of the surfaceome across cancers is likely to grow the small set of currently actionable surface targets. Together, public data and field level commentary support a move from opportunistic target picking toward systematic, population informed discovery. NCBI+1 TALK Decoding the Cell Surface to Accelerate Discovery Program design, trial execution, and measurement Programs that embrace surface resolved selection can operationalize three habits that improve downstream signal. First, connect discovery assays to clinical screening early. If discovery uses a fractionated surface proteomics threshold, define the clinical assay that will mirror that readout, for example an IHC score or a validated flow protocol, and harmonize cutoffs before first patient in. Second, pre specify enrichment rules in the protocol. Enrolling only the top percentiles of surface expressors may seem restrictive, but it increases the chance of a pharmacodynamic signal that validates the mechanism and informs dose expansion. Third, measure what happens when selection criteria are varied in sensitivity analyses, and share these details in publications and regulatory interactions. This transparency builds cumulative knowledge that benefits the field and informs the next iteration of thresholds. There is also value in linking surface metrics to pharmacology. If a surface antigen is abundant but shows slow internalization, a payload with a bystander effect may be preferable to a payload that requires rapid lysosomal routing. If surface expression is heterogeneous at the lesion level, a radionuclide therapy that can exploit crossfire may offer advantages over a conventional ADC. These are not general rules, they are examples of how a surface resolved view can shape the choice of modality and payload in a way that reflects the physical constraints of the target. Where Champions Oncology contributes Champions Oncology generates surface resolved datasets from low passage, clinically relevant tumor models and integrates them with deep multi omic profiles and in vivo pharmacology. This enables teams to quantify true surface positivity, to set and test actionable thresholds, and to understand prevalence by indication before committing to costly trials. The same datasets support orthogonal validation and method standardization so that discovery assays translate into clinical screening. The approach is designed to be non promotional and data first, the objective is to clarify risk and to help programs make better decisions earlier.
Beyond Single Omic Biomarkers: How Proteogenomic ML Reveals Therapy Vulnerabilities
Why a functional view changes predictions Genomics and transcriptomics remain foundational for precision oncology, but they do not fully represent the functional state that determines how tumors respond to therapy. Proteins and phosphoproteins capture activity at the level where drugs actually engage, for example receptor density and localization, complex assembly, and pathway signaling. That distinction is not academic. In a 2024 pan-cancer analysis from the Clinical Proteomic Tumor Analysis Consortium (CPTAC), investigators integrated proteogenomic data from 1,043 patients across 10 tumor types, surveyed 2,863 druggable proteins, and quantified biological factors that weaken mRNA to protein correlation, making the case for models that learn directly from protein and phosphoprotein context rather than inferring from transcripts alone. Cell From data to models that travel The practical bottleneck has been access to harmonized, well-annotated cohorts that support training, testing, and independent validation. In August 2023, the National Cancer Institute announced a standardized pan-cancer proteogenomic dataset that aligns genomics, proteomics, imaging, and clinical data for more than 1,000 tumors across 10 cancer types, explicitly to enable reproducible discovery and model benchmarking. The Proteomic Data Commons (PDC) now serves these resources in a way that supports programmatic access and cross-study comparisons, a requirement if machine-learning outputs are going to generalize beyond a single study. National Cancer Institute+1 Solving the Surfaceome Problem What the evidence shows when proteins are included Two Cell papers from 2024 illustrate why adding protein-level information changes conclusions. An immune-landscape analysis derived distinct immune subtypes by integrating genomic, epigenomic, transcriptomic, and proteomic features and connected oncogenic drivers to downstream protein states that influence immune surveillance and evasion. A companion pan-cancer study expanded the landscape of therapeutic opportunities by evaluating thousands of druggable proteins across tissues and documenting where mRNA is a poor proxy for protein, especially in pathways relevant to therapy response. Together, they show that multi-omic modeling, including protein and phosphoprotein features, improves biological interpretability and exposes actionable biology that single-omic approaches overlook. Cell+1 A broader signal from the field The trend is not confined to CPTAC. The Pan-Cancer Proteome Atlas (TPCPA), published in Cancer Cell in 2025, quantified 9,670 proteins across 999 primary tumors representing 22 cancer types using DIA-MS. The atlas offers a tissue-based substrate for target nomination, biomarker discovery, and external validation, and has been highlighted in the trade press for its global availability and immediate relevance to oncology research. Such atlases are valuable because they capture proteomic variability directly in clinical material, not only in cell lines, providing realistic distributions for features that ML models attempt to learn. Cell+2PubMed+2 Why proteogenomic ML improves prediction Integrating proteins and phosphoproteins adds information that is both mechanistic and measurable. First, pathway activity is reflected in phosphorylation states, which function as on–off or rheostat-like controls for signaling. Second, receptor exposure and complex formation at the protein level determine whether a therapy can bind or disrupt a process. Third, protein degradation and post-translational regulation often decouple mRNA abundance from target availability, which explains why transcript-only biomarkers can fail at the bedside. When these features are engineered into models, performance gains are not just numeric; they tend to be more interpretable, mapping to drug-actionable pathways and receptors that clinicians recognize. The 2024 CPTAC studies provide concrete examples. Immune subtypes defined by proteogenomic features correlate with differences in antigen presentation, cytokine signaling, and interferon responses, features with obvious translational implications. The survey of druggable proteins shows wide variation in abundance and localization across tumors and details the contexts where transcript and protein diverge, arguing for protein-aware rules when nominating targets or stratifying patients. Cell+1 What good practice looks like in model building There is a growing consensus on practical guardrails. Independent validation across cohorts is essential to avoid overfitting, and the infrastructure now exists to support that step through the PDC and related CPTAC resources. Feature construction should prioritize pathway-level signals that aggregate individual phospho-sites into kinase or pathway activity because these are more stable across cohorts and easier to interpret for clinical decision making. Finally, clinically annotated samples, including treatment history and outcomes, are indispensable if models are expected to inform responder enrichment and mechanism-of-resistance hypotheses rather than only classify molecular subtypes. National Cancer Institute Translational payoffs, with appropriate caution When executed with these guardrails, proteogenomic ML offers tangible benefits. Programs can generate earlier responder and non-responder hypotheses and test them prospectively in preclinical systems before committing costly clinical designs. Resistance pathways inferred from phospho-proteomic features can motivate combination strategies, for example pairing an antibody-drug conjugate with a kinase inhibitor when signaling indicates a plausible escape route. Educational content from ASCO has emphasized the centrality of quantitative thresholds and validated assays for patient selection, particularly for ADCs where surface accessibility and abundance determine benefit. The lesson is consistent across modalities. Predictive features must be connected to assays that can be deployed consistently in trials, and thresholds should be defined in a way that anticipates real-world variability. PubMed Where limitations still matter Several limitations deserve explicit mention. Proteomic and phospho-proteomic data remain technically variable across platforms and laboratories. Although CPTAC and PDC mitigate this through standardization, modelers should evaluate batch effects and apply normalization strategies suited to proteomic data. Coverage of kinase–substrate relationships and post-translational networks is incomplete, which constrains inference. Tumor heterogeneity adds another layer, particularly when bulk tissue averages mask subclonal or microenvironmental signals. These caveats do not negate the value of proteogenomic ML, but they do argue for conservative claims, orthogonal validation, and a bias toward features that can be measured reproducibly in clinical settings. TALK Decoding the Cell Surface to Accelerate Discovery Implications for trial design and portfolio focus The immediate implication is a more disciplined approach to enrichment. If protein-level features identify a subgroup with plausible sensitivity, early designs can incorporate eligibility criteria and stratification based on validated assays rather than exploratory cutpoints. Conversely, if pathway-level features suggest multiple escape routes, it may be more efficient to prioritize combinations earlier instead of iterating single-agent studies. At a portfolio level, proteogenomic evidence can help prioritize programs with a mechanistic rationale supported by functional data, not only by mutation prevalence or gene expression. How Champions Oncology contributes Champions Oncology builds models on tumor-derived systems that preserve patient biology and heterogeneity. Our datasets link genomics and transcriptomics with proteomics, phospho-proteomics, and cell surface proteomics, and they are annotated with pharmacologic phenotypes. This combination supports models that tie features to functional biology and drug accessibility, making it possible to move from correlation to causal relation and from causal relation to druggable targets.

Feature selection in RNA-seq and proteomics with MADVAR in R
High dimensional omics datasets often include many features that contribute little to downstream analysis. This can blur structure in unsupervised tasks, slow computation, and complicate model training. The MADVAR study introduces two simple, data driven procedures that set feature selection thresholds from the distribution of the data itself, rather than relying on fixed heuristics. The first procedure, madvar, computes a variance cutoff using the median plus a multiple of the median absolute deviation. The second, intersect Distributions, fits a two component Gaussian mixture to the variance or another continuous score, and uses the intersection point between components as the cutoff. Both methods are implemented in an R package and were evaluated across public datasets that include TCGA gene expression, GTEx proteomics, and CPTAC phosphoproteomics. The paper reports improvements in unsupervised clustering quality and competitive supervised performance with fewer features, while keeping runtime and memory use modest. What the Paper Tested The benchmarking examined unsupervised structure and supervised classification. For unsupervised analysis, the authors applied filtering and then assessed cluster quality with connectivity, the Dunn index, and the Biological Homogeneity Index. Across datasets, the variance based approaches produced tighter or more homogeneous clusters on these metrics. For supervised analysis, they trained random forest models with repeated runs. Both approaches produced low out of bag error rates. Retaining more features sometimes improved accuracy, but MADVAR often matched the mixture based approach while selecting fewer features. The paper also documents practical defaults, such as Ward.D for hierarchical clustering with Euclidean distance, and explains how to pass either a raw matrix or a precomputed variance vector into the functions. Source code and documentation are available on GitHub. When These Methods are a Good Fit These procedures are particularly well suited for rapid, large-scale preprocessing, when analyses require a quick, efficient, and transparent approach to feature selection prior to dimensionality reduction, clustering, or model fitting. They also integrate naturally into interpretability-oriented pipelines, since thresholds based on medians or mixture-model intersections are simple to explain and justify to collaborators. Because the logic operates on continuous feature scores, the same framework can be applied seamlessly to any quantitative data type that can be summarized by variability Things to keep in mind. Variance is a proxy for informativeness, not a guarantee. Low variance does not always mean a feature is uninformative. Some biomarkers remain stable yet become predictive in combination with others. If domain knowledge indicates a feature should be preserved, the package allows must keep lists. The mixture based method assumes the variance distribution resembles a two-component mixture. If the fit is poor, the intersection may not be meaningful, so density plots are worth inspecting before adopting the cutoff. Downstream metric choice also matters. Gains in the Biological Homogeneity Index or Dunn index describe cluster characteristics, which may not translate to improvements on other endpoints such as survival modeling or dose response prediction. Finally, supervised performance can depend on class imbalance and sample size. If your data are skewed, tune the learner and validate with a scheme that reflects your use case. Read the Full Paper How to Apply, a Straightforward Workflow A practical workflow starts by exploring the variance distribution. Plot the density (using the madvar flag `plot_density = TRUE`), confirm whether there is a near zero peak, and decide whether a MAD based threshold or a two-component mixture are appropriate. Set a conservative first pass using the default MAD multiplier (`mads = 2`) and adjust if another stringency level is preferred, or, if you prefer the mixture approach, verify the intersection visually before you commit to the cutoff. Preserve domain critical features by whitelisting known markers or controls that should not be dropped. Re run the planned clustering or model fitting, compare structure and error rates before and after filtering, and record any change in feature counts and compute time so the impact is transparent to collaborators. Reproducibility and availability The R implementation and documentation are available on GitHub, as referenced in the paper. The evaluations draw on TCGA gene expression, GTEx proteomics, and CPTAC phosphoproteomics, with figures that show density plots, clustering metrics, and classification results. The article appears as an Application Note in Bioinformatics Advances and is available for open access. Summary MADVAR provides two transparent, variance-based rules for feature selection, enabling the removal of near invariant features from large omics matrices. In the reported benchmarks, these procedures improved or maintained clustering quality and supervised accuracy while substantially reducing feature counts and computational load. The approach is easy to inspect, easy to explain, and simple to integrate into existing R workflows. As with any filter, final value depends on the analysis goal, so it is worth validating its effects on the endpoints that matter for your study. Silberberg G. “MADVAR, a lightweight, data driven tool for automated feature selection in omics data.” Bioinformatics Advances 2025, vbaf211. doi, 10.1093/bioadv/vbaf211. Explore our Data Ecosystem

Choosing the Right CDX Models: Speed, Scientific Value, and Translational Relevance
In oncology drug development, cell line-derived xenograft (CDX) models remain one of the cornerstone in vivo tools for evaluating new therapies in the preclinical ecosystem. They offer a balance of biological relevance, reproducibility, and speed that makes them ideal for early phase hypothesis testing. Nevertheless, while CDX models are indispensable for initial target screening and validation, they also have well-understood limitations that can upset downstream progress if not properly accounted for. As drug developers increasingly rely on CDX-based systems to screen and prioritize compounds before moving into clinically relevant, albeit more expensive, patient-derived xenografts (PDX), the need for appropriate model selection has become critical. The right CDX program can deliver early translational clarity and strategic focus, while the wrong one can generate noise that obscures the true efficacy of an otherwise promising therapeutic, potentially derailing its continued development. Balancing Speed and Fidelity CDX studies are often designed for speed. They enable large-scale screening of novel agents, as both mono- and combination therapies, and dosing regimens in a fraction of the time and cost required for PDX studies. This makes them especially valuable at the earliest stages of decision-making, where timelines are incredibly compressed and attrition risk is high. But that same speed exacts a cost. Many CDX models originate from long established cell lines that have been maintained in culture for decades. Over time, these lines lose virtually all the critical genomic and cellular heterogeneity, stromal interactions, and microenvironmental complexity characteristic of a bona fide tumor. Whilst they grow predictably, the biological features they retain may no longer reflect that of tumors seen in clinical patients. However, the key is not to abandon CDX models, but to recognize where they fit in the development pipeline and to acknowledge and mitigate the limitations they have. Used prudently, CDX models are an efficient and scientifically powerful system to rank compounds, explore mechanisms, and refine hypotheses before moving into PDX for deeper translational validation. Defining a High Value CDX Model The quality of a CDX model is defined by its biological fidelity and characterization depth. Models derived from contemporary, clinically annotated cell lines are more likely to capture the genomic and phenotypic diversity relevant to modern oncology therapeutics. When models reflect the intrinsic complexity of current patient populations, such as the plethora of KRAS G12 mutations or the legion mechanisms by which EGFR becomes hyperactivated, they offer drug developers meaningful results and mechanism-linked insights that can inform clinical strategy. Biological and “omic” characterization matters as much, if not more, than cellular/tissue origin in CDX models, particularly as clinical oncology continues to diverge from tissue-based therapy to therapeutic roadmaps grounded in the molecular features of each patient tumor. A high value CDX model is supported by multi-omic profiling that includes genomic, transcriptomic, and proteomic annotation. This data allows researchers to interpret observed drug effects through the lens of pathway activation, resistance mechanisms, and biomarker expression. In contrast, models lacking such characterization risk generating results that are descriptive rather than explanatory. Drive Proof of Concept and Target Validation with Champions’ CDX Models Mechanistic Clarity Through Multi-Omic Integration The rise of precision oncology has shown that pharmacology and data science are inextricably linked. Researchers now expect and rely on preclinical models to yield mechanistic understanding and insight, not just tumor regression rates. Integrating omic datasets into CDX studies transforms them from mere screening tools into translational resources capable of generating biological comprehension and preclinical direction. For instance, RNA sequencing of treated and untreated xenografts can reveal transcriptional signatures underpinning positive response outcomes, potentially allowing clinical partitioning of patients likely to receive the most benefit from a therapy. As another example, phosphoproteomic profiling can identify compensatory signaling cascades mediating adaptive drug resistance, permitting de novo combination therapies to be trialed to preempt such resistance before it evolves. This approach enables drug developers to anticipate how tumors might evade inhibition, long before clinical exposure. Moreover, omic integration provides a framework for cross platform alignment. Data from CDX models can be mapped against PDX datasets, public repositories, or patient trial cohorts, accelerating the feedback loop between preclinical findings and clinical validation. Modeling Resistance and Tumor Evolution One of the most powerful applications of CDX technologies is in modeling acquired resistance. By exposing tumor-bearing mice to sustained drug pressure, scientists can select for resistant clones that mimic clinical relapse. Comparative molecular profiling between parental and resistant CDX lines may illuminate the pathways that drive therapeutic escape, whether through secondary genomic changes, activation of bypass signaling cascades, or metabolic rewiring. This approach supports the rational design of next-generation inhibitors or combination strategies aimed at delaying or overcoming resistance. It also informs biomarker development by revealing the early molecular changes that forecast reduced drug sensitivity, enabling the design of clinical trials with built-in resistance monitoring. Bridging the Gap to PDX Despite their wide-ranging utility and flexibility as experimental tools, CDX models are at best a starting technology in the developmental pipeline. Translationally-minded organizations deploy CDX models as a filter for promising candidates in an experimental continuum that leads naturally into PDX. PDX models, established directly from patient tumors, preserve the architecture, stromal components, and molecular heterogeneity of the original cancer. They capture biological features that CDXs inherently lack, including contextualized immune responses in humanized systems, evolution and expansion of tumor sub-clones, and the influence of the microenvironment on cancer progression. For these reasons, PDX validation remains an important next step, if not a critical one, once a compound demonstrates clear activity in CDX. The most efficient development pipelines are those where CDX and PDX models are in biological alignment, where both originate from well characterized sources and overlap genomically and/or phenotypically. In this context, compound evaluation flows smoothly from screening to translational evaluation. A consistent molecular linkage between model systems strengthens the predictive bridge and ensures that early results translate more faithfully into clinical outcomes. Speed Without Sacrificing Relevance The enduring appeal of CDX model systems lies in the speed with which large quantities of data can be generated to reinforce or oppose development of individual drug compounds, or indeed entire drug programs. Studies can be initiated quickly, with timelines to results measured in weeks rather than months. Moreover, CDX models can support the simultaneous exploration of multiple therapeutic hypotheses. Where delays mean patients remain beset with therapeutic inadequacies and can cost developers millions in lost opportunity, this speed is a major competitive advantage. But whilst speed is a necessary component of drug development, it is insufficient for success. The most effective drug pipelines are designed with translational intent incorporated into CDX programs from the outset. CDX model selection is based on mechanistic alignment derived from deep omics comprehensive characterization, and the clinical transition to PDX model systems is a crucial element of the sequence rather than an unintegrated effort. This approach ensures CDX models are deployed as value-enhancing study tools,leveraging the efficiency of CDXs to inform smarter, faster progression into the PDX models best approximating patient clinical features and responses. Designing a Modern CDX Strategy For biotechnology and pharmaceutical teams, a modern CDX strategy balances three principles:speed, depth, and connectivity. Speed means using CDX models to quickly triage candidate molecules, confirm on-target effects, and eliminate ineffective compounds prior to larger resource investment. Depth refers to multi-layered omics characterization to uncover mechanistic drivers of response or resistance. And finally, connectivity means designing CDX studies with the downstream transition to PDX models in mind, ensuring molecular alignment and continuity between the different systems. When these principles are applied judiciously, CDX models become a strategic asset that accelerates development timelines without compromising scientific integrity. The Future of Translational Modeling As oncology continues to evolve toward precision medicine, the most impactful preclinical programs will be those that connect fast data generation with deep omics characterization, using rapid CDX screening to guide more complex, patient-relevant studies. Emerging approaches such as multi-omic analytics, AI-driven model selection, and ex vivo/organoid validation are expanding how CDX data can inform clinical decision-making. Taken in concert, these all suggest a future where the value of a model is defined not only by how fast it can be employed to produce data, but also by how that same data can be used to map the necessary downstream steps to ensure successful drug development and patient application. Choosing the Right Partner In an increasingly competitive preclinical landscape, the distinction between a vendor and a scientific partner has never been clearer. The most valuable CDX programs combine biological relevance, data transparency, and translational foresight. When evaluating potential collaborators, sponsors should ask not only what models are available but how those models were developed, characterized, and validated. The answers will reveal whether a platform can deliver more than results, whether it can deliver understanding. Cell Line Select Tool A smarter way to explore Champions’ cell line models.
Modeling next generation AR pathway inhibitors in prostate cancer
When Inhibition Isn’t Enough: How Dual-Mechanism AR Degraders Are Redefining Resistance Modeling For over a decade, androgen receptor (AR) pathway inhibitors such as enzalutamide and abiraterone have formed the foundation of treatment for advanced prostate cancer. Yet the same story continues to unfold patients initially respond, then relapse. Despite continued suppression of androgen signaling, tumors adapt reactivating the AR axis through overexpression, gene amplification, or ligand-binding domain mutations. What follows is a return of disease activity that current inhibitors can no longer control. This persistent pattern highlights a central truth in oncology drug development: inhibition alone is not enough. Over time, cells find ways to reengage the same signaling pathways that were once silenced. A new class of therapies is changing that. Bristol Myers Squibb recently published a landmark study in Clinical Cancer Research describing BMS-986365, a first-in-class dual AR degrader and antagonist. By combining proteasomal degradation with receptor antagonism, BMS-986365 achieves a more profound and sustained blockade of AR activity than traditional inhibitors. The drug not only shuts down signaling but also removes the receptor protein responsible for resistance and relapse. Learn more about our Prostate Models Champions Oncology’s patient-derived xenograft (PDX) models CTG-2440 and CTG-2441 were instrumental in this discovery. These models were derived from the same patient before and after abiraterone therapy, creating a unique, clinically matched system for studying adaptive resistance. In both models, BMS-986365 was tested alongside enzalutamide to evaluate its impact on AR signaling in tumors that had already progressed on prior therapy. While the model developed before the patient received abiraterone showed modest increased sensitivity to BMS-986365 compared to Enzalutamide, BMS-986365 outperformed Enzalutamide when administered to the model developed after the patient progressed under abiraterone. Both drugs increased AR mRNA expression in treated tumors, q a typical feedback response to pathway inhibition but only BMS-986365, by virtue of its degrader activity, maintained low AR protein levels and continued to suppress AR target gene activity. This finding underscores the importance of degradation: while transcriptional upregulation persisted, the protein was degraded before it could restore signaling. The biological feedback loop driving adaptive resistance was disrupted during treatment. These insights speak to a broader principle in translational research: resistance is dynamic, not static. It doesn’t exist as a single genetic event, but as a continuum of cellular responses that evolve under therapeutic pressure. Capturing this dynamic behavior requires preclinical models that replicate the complexity of human disease including treatment history, adaptive feedback mechanisms, and the heterogeneous signaling environments of late-line tumors. For drug developers, this means that testing next-generation degraders and dual-mechanism agents cannot rely on traditional cell lines or simplified, outdated in vivo models. The field now demands functional, clinically grounded resistance models that measure more than endpoint response they must reveal how and why tumors adapt, and how new modalities can overcome that adaptation. Champions Oncology’s portfolio of pretreated and resistance-matched PDX models was built precisely for this purpose. By recreating clinical resistance within the same biological context in which it arises, these models provide a high-fidelity platform to evaluate degrader pharmacodynamics, durability of response, and combination potential. The BMS-986365 study offers a clear demonstration of their value: real-world resistance biology, translated into preclinical discovery that informed a novel therapeutic strategy. The success of BMS-986365 represents more than a promising drug—it marks a shift in how we define preclinical relevance. By pairing innovative therapeutics with equally advanced resistance models, the field is beginning to close the translational gap that has long limited success in late-stage prostate cancer. Discover Pretreated and Resistance-Matched PDX Models
